Ace+Method+Development+Kits
Catalog Number:
(76457-536)
Supplier:
VWR International
Description:
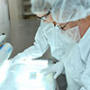
Wide variety of VWR® student dissection kits that suit educational needs.
Catalog Number:
(77440-326)
Supplier:
Bioss
Description:
Calcium- and diacylglycerol-independent serine/threonine-protein kinase that functions in phosphatidylinositol 3-kinase (PI3K) pathway and mitogen-activated protein (MAP) kinase cascade, and is involved in NF-kappa-B activation, mitogenic signaling, cell proliferation, cell polarity, inflammatory response and maintenance of long-term potentiation (LTP). Upon lipopolysaccharide (LPS) treatment in macrophages, or following mitogenic stimuli, functions downstream of PI3K to activate MAP2K1/MEK1-MAPK1/ERK2 signaling cascade independently of RAF1 activation. Required for insulin-dependent activation of AKT3, but may function as an adapter rather than a direct activator. Upon insulin treatment may act as a downstream effector of PI3K and contribute to the activation of translocation of the glucose transporter SLC2A4/GLUT4 and subsequent glucose transport in adipocytes. In EGF-induced cells, binds and activates MAP2K5/MEK5-MAPK7/ERK5 independently of its kinase activity and can activate JUN promoter through MEF2C. Through binding with SQSTM1/p62, functions in interleukin-1 signaling and activation of NF-kappa-B with the specific adapters RIPK1 and TRAF6. Participates in TNF-dependent transactivation of NF-kappa-B by phosphorylating and activating IKBKB kinase, which in turn leads to the degradation of NF-kappa-B inhibitors. In migrating astrocytes, forms a cytoplasmic complex with PARD6A and is recruited by CDC42 to function in the establishment of cell polarity along with the microtubule motor and dynein. In association with FEZ1, stimulates neuronal differentiation in PC12 cells. In the inflammatory response, is required for the T-helper 2 (Th2) differentiation process, including interleukin production, efficient activation of JAK1 and the subsequent phosphorylation and nuclear translocation of STAT6. May be involved in development of allergic airway inflammation (asthma), a process dependent on Th2 immune response. In the NF-kappa-B-mediated inflammatory response, can relieve SETD6-dependent repression of NF-kappa-B target genes by phosphorylating the RELA subunit at 'Ser-311'. Necessary and sufficient for LTP maintenance in hippocampal CA1 pyramidal cells. In vein endothelial cells treated with the oxidant peroxynitrite, phosphorylates STK11 leading to nuclear export of STK11, subsequent inhibition of PI3K/Akt signaling, and increased apoptosis. Phosphorylates VAMP2 in vitro (PubMed:17313651).
Catalog Number:
(10109-534)
Supplier:
Prosci
Description:
Kynureninase is a pyridoxal-5'-phosphate (pyridoxal-P) dependent enzyme that catalyzes the cleavage of L-kynurenine and L-3-hydroxykynurenine into anthranilic and 3-hydroxyanthranilic acids, respectively. Kynureninase is involved in the biosynthesis of NAD cofactors from tryptophan through the kynurenine pathway.Kynureninase is a pyridoxal-5'-phosphate (pyridoxal-P) dependent enzyme that catalyzes the cleavage of L-kynurenine and L-3-hydroxykynurenine into anthranilic and 3-hydroxyanthranilic acids, respectively. Kynureninase is involved in the biosynthesis of NAD cofactors from tryptophan through the kynurenine pathway. Two transcript variants encoding different isoforms have been found for this gene.Kynureninase is a pyridoxal-5'-phosphate (pyridoxal-P) dependent enzyme that catalyzes the cleavage of L-kynurenine and L-3-hydroxykynurenine into anthranilic and 3-hydroxyanthranilic acids, respectively. Kynureninase is involved in the biosynthesis of NAD cofactors from tryptophan through the kynurenine pathway. Two transcript variants encoding different isoforms have been found for this gene.
Catalog Number:
(10111-196)
Supplier:
Prosci
Description:
The 'SNARE hypothesis' is a model explaining the process of docking and fusion of vesicles to their target membranes. According to this model, membrane proteins from the vesicle (v-SNAREs) and proteins from the target membrane (t-SNAREs) govern the specificity of vesicle targeting and docking through mutual recognition. Once the 2 classes of SNAREs bind to each other, they form a complex that recruits the general elements of the fusion apparatus, namely NSF (N-ethylmaleimide-sensitive factor) and SNAPs (soluble NSF-attachment proteins), to the site of membrane fusion, thereby forming the 20S fusion complex. Alpha- and gamma-SNAP are found in a wide range of tissues and act synergistically in intra-Golgi transport. The sequence of the predicted 295-amino acid human protein encoded by NAPA shares 37%, 60%, and 67% identity with the sequences of yeast, Drosophila, and squid alpha-SNAP, respectively.
Catalog Number:
(10108-796)
Supplier:
Prosci
Description:
DNASE2B shares considerable sequence similarity to, and is structurally related to DNase II. The latter is a well characterized endonuclease that catalyzes DNA hydrolysis in the absence of divalent cations at acidic pH. Unlike DNase II which is ubiquitously expressed, expression of this protein is restricted to the salivary gland and lungs.The protein encoded by this gene shares considerable sequence similarity to, and is structurally related to DNase II. The latter is a well characterized endonuclease that catalyzes DNA hydrolysis in the absence of divalent cations at acidic pH. Unlike DNase II which is ubiquitously expressed, expression of this gene product is restricted to the salivary gland and lungs. The gene has been localized to chromosome 1p22.3 adjacent (and in opposite orientation) to the uricase pseudogene. Two transcript variants encoding different isoforms have been described for this gene.
Catalog Number:
(10101-920)
Supplier:
Prosci
Description:
GJA4 is a member of the connexin family. The protein is a component of gap junctions, which are composed of arrays of intercellular channels that provide a route for the diffusion of low molecular weight materials from cell to cell. Mutations in this gene have been associated with atherosclerosis and a higher risk of myocardial infarction.This gene encodes a member of the connexin gene family. The encoded protein is a component of gap junctions, which are composed of arrays of intercellular channels that provide a route for the diffusion of low molecular weight materials from cell to cell. Mutations in this gene have been associated with atherosclerosis and a higher risk of myocardial infarction. Publication Note: This RefSeq record includes a subset of the publications that are available for this gene. Please see the Entrez Gene record to access additional publications.
Catalog Number:
(10111-636)
Supplier:
Prosci
Description:
MGAT2 is a Golgi enzyme catalyzing an essential step in the conversion of oligomannose to complex N-glycans. The enzyme has the typical glycosyltransferase domains: a short N-terminal cytoplasmic domain, a hydrophobic non-cleavable signal-anchor domain, and a C-terminal catalytic domain. Mutations in its gene may lead to carbohydrate-deficient glycoprotein syndrome, type II.The product of this gene is a Golgi enzyme catalyzing an essential step in the conversion of oligomannose to complex N-glycans. The enzyme has the typical glycosyltransferase domains: a short N-terminal cytoplasmic domain, a hydrophobic non-cleavable signal-anchor domain, and a C-terminal catalytic domain. Mutations in this gene may lead to carbohydrate-deficient glycoprotein syndrome, type II. Two transcript variants encoding the same protein have been identified for this gene.
Catalog Number:
(10110-554)
Supplier:
Prosci
Description:
The structure of MAS1 indicates that it belongs to the class of receptors that are coupled to GTP-binding proteins and share a conserved structural motif, which is described as a '7-transmembrane segment' following the prediction that these hydrophobic segments form membrane-spanning alpha-helices. The MAS1 protein may be a receptor that, when activated, modulates a critical component in a growth-regulating pathway to bring about oncogenic effects.The structure of the MAS1 product indicates that it belongs to the class of receptors that are coupled to GTP-binding proteins and share a conserved structural motif, which is described as a '7-transmembrane segment' following the prediction that these hydrophobic segments form membrane-spanning alpha-helices. The MAS1 protein may be a receptor that, when activated, modulates a critical component in a growth-regulating pathway to bring about oncogenic effects.
Catalog Number:
(10102-298)
Supplier:
Prosci
Description:
SIRT5 is included in class III of the sirtuin family which ischaracterized by a sirtuin core domain. Human sirtuins may function as intracellular regulatory proteins with mono-ADP-ribosyltransferase activity. This gene encodes a member of the sirtuin family of proteins, homologs to the yeast Sir2 protein. Members of the sirtuin family are characterized by a sirtuin core domain and grouped into four classes. The functions of human sirtuins have not yet been determined; however, yeast sirtuin proteins are known to regulate epigenetic gene silencing and suppress recombination of rDNA. Studies suggest that the human sirtuins may function as intracellular regulatory proteins with mono-ADP-ribosyltransferase activity. The protein encoded by this gene is included in class III of the sirtuin family. Alternative splicing of this gene results in two transcript variants.
Catalog Number:
(10106-484)
Supplier:
Prosci
Description:
The protein encoded by MCM4 is one of the highly conserved mini-chromosome maintenance proteins (MCM) that are essential for the initiation of eukaryotic genome replication. The hexameric protein complex formed by MCM proteins is a key component of the pre-replication complex (pre_RC) and may be involved in the formation of replication forks and in the recruitment of other DNA replication related proteins. The MCM complex consisting of this protein and MCM2, 6 and 7 proteins possesses DNA helicase activity, and may act as a DNA unwinding enzyme. The phosphorylation of this protein by CDC2 kinase reduces the DNA helicase activity and chromatin binding of the MCM complex. The MCM4 gene is mapped to a region on the chromosome 8 head-to-head next to the PRKDC/DNA-PK, a DNA-activated protein kinase involved in the repair of DNA double-strand breaks.
Catalog Number:
(10104-462)
Supplier:
Prosci
Description:
The first mRNA transcript isolated for C2orf3 gene was part of an artificial chimera derived from two distinct gene transcripts and a primer used in the cloning process (see Genbank accession M29204). C2orf3 belongs to the GCF family. It is a factor that represses transcription. It binds to the GC-rich sequences (5'-GCGGGGC-3') present in the epidermal growth factor receptor, beta-actin, and calcium-dependent protease promoters.The first mRNA transcript isolated for this gene was part of an artificial chimera derived from two distinct gene transcripts and a primer used in the cloning process (see Genbank accession M29204). A positively charged amino terminus present only in the chimera was determined to bind GC-rich DNA, thus mistakenly thought to identify a transcription factor gene.
Catalog Number:
(10111-368)
Supplier:
Prosci
Description:
E2F5 is a member of the E2F family of transcription factors. The E2F family plays a crucial role in the control of cell cycle and action of tumor suppressor proteins and is also a target of the transforming proteins of small DNA tumor viruses. The E2F proteins contain several evolutionally conserved domains found in most members of the family. These domains include a DNA binding domain, a dimerization domain which determines interaction with the differentiation regulated transcription factor proteins (DP), a transactivation domain enriched in acidic amino acids, and a tumor suppressor protein association domain which is embedded within the transactivation domain. This protein is differentially phosphorylated and is expressed in a wide variety of human tissues. It is more homologous to E2F4, a family member, than to other members. Both this protein and E2F4 interact with tumor suppressor proteins p130 and p107, but not with pRB.
Catalog Number:
(10106-468)
Supplier:
Prosci
Description:
Gap junctions were first characterized by electron microscopy as regionally specialized structures on plasma membranes of contacting adherent cells. These structures were shown to consist of cell-to-cell channels. Proteins, called connexins, purified from fractions of enriched gap junctions from different tissues differ. The connexins are designated by their molecular mass. Another system of nomenclature divides gap junction proteins into 2 categories, alpha and beta, according to sequence similarities at the nucleotide and amino acid levels. For example, CX43 (MIM 121014) is designated alpha-1 gap junction protein, whereas CX32 (GJB1; MIM 304040) and CX26 are called beta-1 and beta-2 gap junction proteins, respectively. This nomenclature emphasizes that CX32 and CX26 are more homologous to each other than either of them is to CX43.
Catalog Number:
(10106-762)
Supplier:
Prosci
Description:
L-type calcium channels are composed of five subunits. CACNG4 represents one of these subunits, gamma, and is one of several gamma subunit proteins. It is an integral membrane protein that is thought to stabilize the calcium channel in an inactive (closed) state. This gene is a member of the neuronal calcium channel gamma subunit gene subfamily of the PMP-22/EMP/MP20 family.L-type calcium channels are composed of five subunits. The protein encoded by this gene represents one of these subunits, gamma, and is one of several gamma subunit proteins. It is an integral membrane protein that is thought to stabilize the calcium channel in an inactive (closed) state. This gene is a member of the neuronal calcium channel gamma subunit gene subfamily of the PMP-22/EMP/MP20 family and is located in a cluster with two similar gamma subunit-encoding genes.
Catalog Number:
(10102-392)
Supplier:
Prosci
Description:
This gene is a paralog of SMN1 gene, which encodes the survival motor neuron protein, mutations in which are cause of autosomal recessive proximal spinal muscular atrophy. SMNDC1 is a nuclear protein that has been identified as a constituent of the spliceosome complex. This gene is differentially expressed, with abundant levels in skeletal muscle, and may share similar cellular function as the SMN1 gene. This gene is a paralog of SMN1 gene, which encodes the survival motor neuron protein, mutations in which are cause of autosomal recessive proximal spinal muscular atrophy. The protein encoded by this gene is a nuclear protein that has been identified as a constituent of the spliceosome complex. This gene is differentially expressed, with abundant levels in skeletal muscle, and may share similar cellular function as the SMN1 gene.
Catalog Number:
(10111-098)
Supplier:
Prosci
Description:
Regulator of G protein signaling (RGS) family members are regulatory molecules that act as GTPase activating proteins (GAPs) for G alpha subunits of heterotrimeric G proteins. RGS proteins are able to deactivate G protein subunits of the Gi alpha, Go alpha and Gq alpha subtypes. They drive G proteins into their inactive GDP-bound forms. Regulator of G protein signaling 10 belongs to this family. All RGS proteins share a conserved 120-amino acid sequence termed the RGS domain. This protein associates specifically with the activated forms of the two related G-protein subunits, G-alphai3 and G-alphaz but fails to interact with the structurally and functionally distinct G-alpha subunits. Regulator of G protein signaling 10 protein is localized in the nucleus. Two transcript variants encoding different isoforms have been found for this gene.
Inquire for Price
Stock for this item is limited, but may be available in a warehouse close to you. Please make sure that you are logged in to the site so that available stock can be displayed. If the
Stock for this item is limited, but may be available in a warehouse close to you. Please make sure that you are logged in to the site so that available stock can be displayed. If the
You must log in to order restricted items. We request that you provide the required business documentation to purchase this product for the first time.
To order chemicals, medical devices, or other restricted products please provide identification that includes your business name and shipping address via email CMD_NA@vwr.com or fax 484.881.5997 referencing your VWR account number . Acceptable forms of identification are:
-Additional Documentation May be needed to purchase this item. A VWR representative will contact you if needed.
This product has been blocked by your organization. Please contact your purchasing department for more information.
The original product is no longer available. The replacement shown is available.
This product is currently unavailable but limited stock may be available in our extended warehouse network. Please call 1-800-932-5000 and a VWR Customer Service Representative will help you.
|
|||||||||